ASLAN reports interim data from atopic dermatitis drug trial
Findings showed that 73.3% of subjects in the eblasakimab arm achieved an EASI-75 response versus 14.3% of those given a placebo.
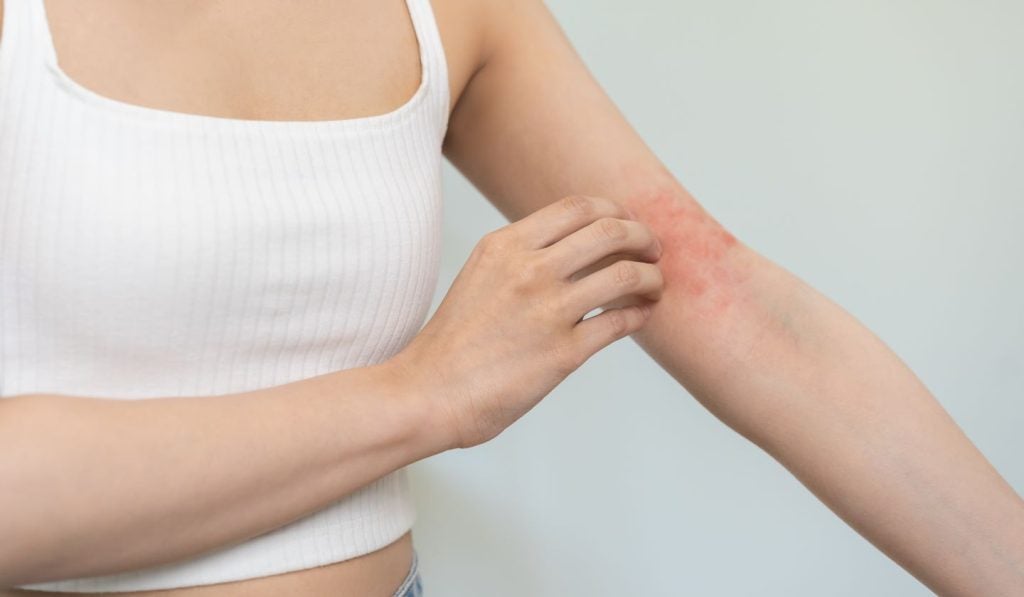
Findings showed that 73.3% of subjects in the eblasakimab arm achieved an EASI-75 response versus 14.3% of those given a placebo.


The observational study will assess the clinical and workflow benefits of using Hyperfine’s Swoop system in detecting amyloid abnormalities in Alzheimer’s patients.
The pharmaceutical industry continues to be a hotbed of patent innovation. Activity is driven by the evolution of new treatment...
Acarix starts clinical workflow study for its AI-powered cardiac diagnostic
Cleerly touts new data for AI cardiovascular software
Moderna’s Phase III Covid-19 vaccine trial meets primary endpoints
Gedeon Richter adopts TransPerfect’s trial management solutions